Visualizing Science 2016: Beautiful Images From Researchers in CNS
As part of an ongoing tradition, this past spring we invited faculty, staff and students in the College of Natural Sciences community to send us images that celebrated the wondrous beauty of science and the scientific process. We were searching for those moments where science and art meld and become one.
Scientific discovery has always been partly a visual pursuit, as researchers attempt to communicate their discoveries in a meaningful manner and explore the topics that hold their fascination. Darwin traced evolutionary trees in his notes, Hooke sketched astronomical and microscopic observations, Franklin made X-ray diffraction images vital to determining the structure of DNA, and Feynman devised diagrams that helped transform theoretical physics, just to name a few.
Now more than ever, scientific visualizations are becoming an integral part of the discovery process. Researchers use 3-D models and data visualizations to reveal patterns hidden within data, to uncover the inner workings of life or to explain the very structure of the universe.
This year the winners were revealed at Science in the Public Square, an event to celebrate the intersection of science and public engagement. Below we feature seven of the most stunning submissions from our scientific community. The first six images were chosen by committee based on their beauty and scientific merit. The final image, our Facebook favorite, was chosen by the public on our Facebook page. The first six images will be displayed in The University of Texas at Austin Tower, Welch Hall, and the Kuehne Physics Mathematics Astronomy Library in Physics, Math & Astronomy Building.
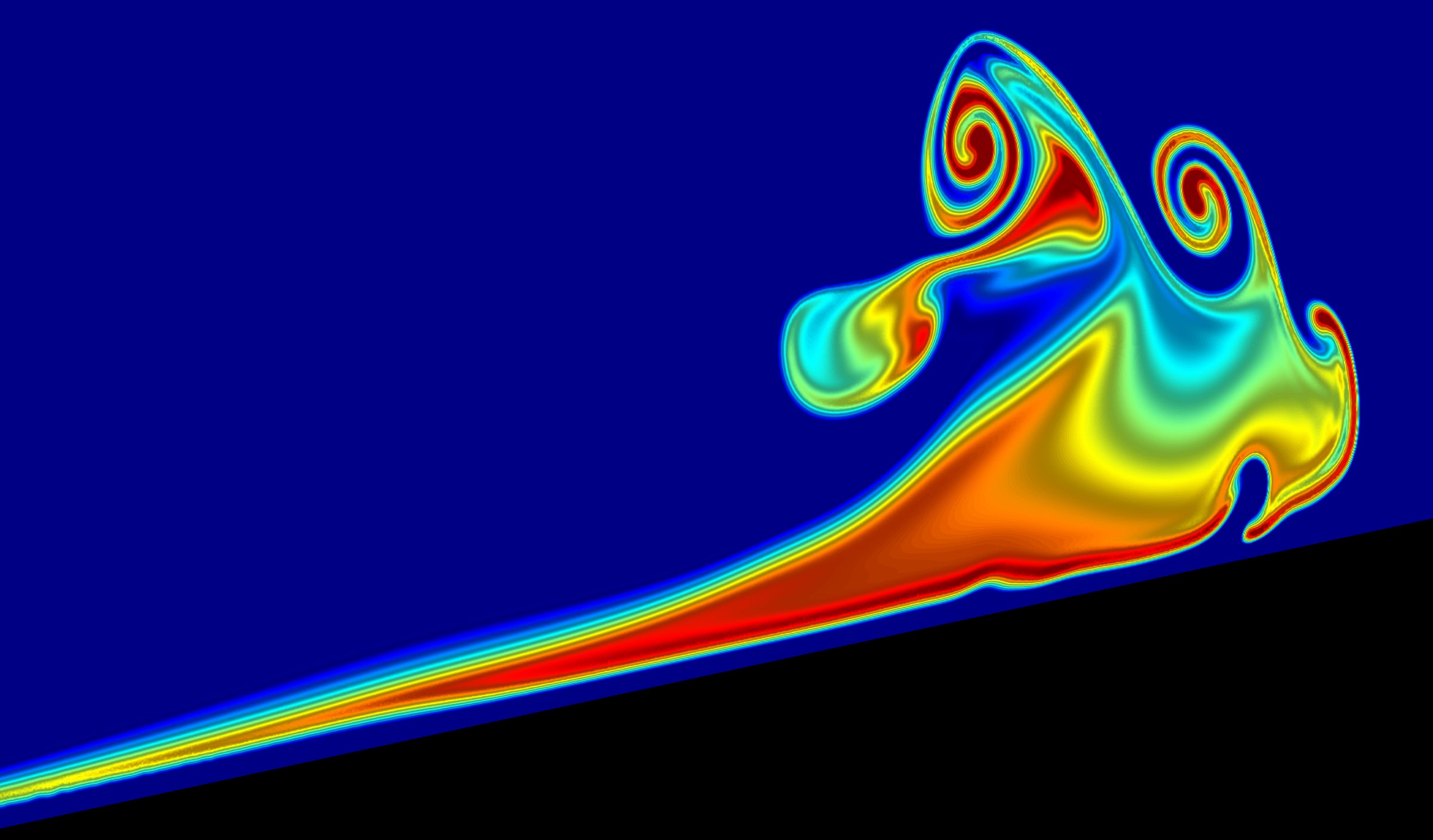
FIRST PLACE
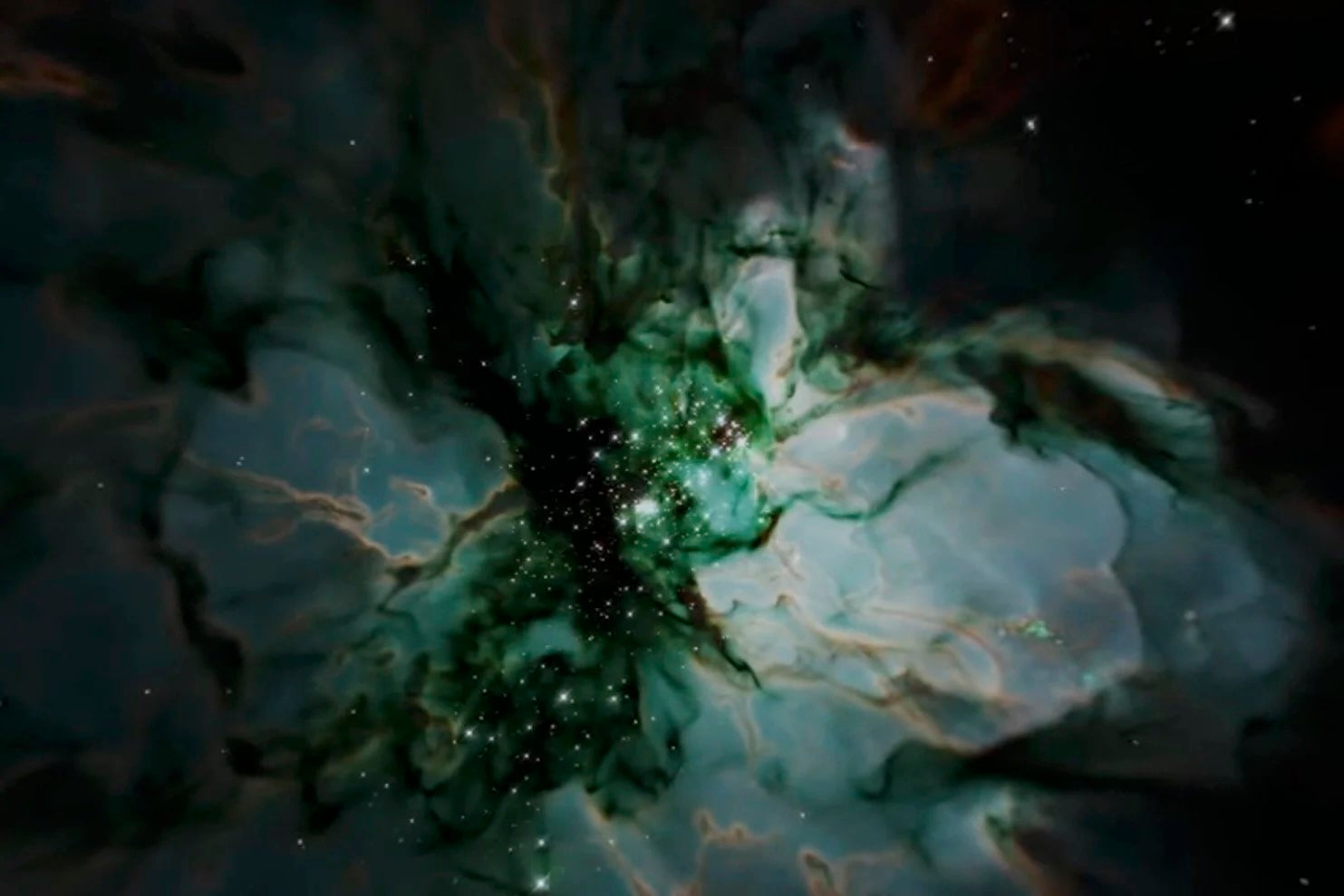
Much like waves crashing on the beach, subsurface waves travel thousands of kilometers across the ocean till the 200-meter-tall waves reach the coastline where they steepen and break. These physicists’ simulations investigate this breaking process, which produces a surge of cold salty water up the slope that has both oceanographic and ecological ramifications. — Michael Allshouse, PhD, Postdoctoral Researcher in the Department of Physics’ Center for Nonlinear Dynamics.
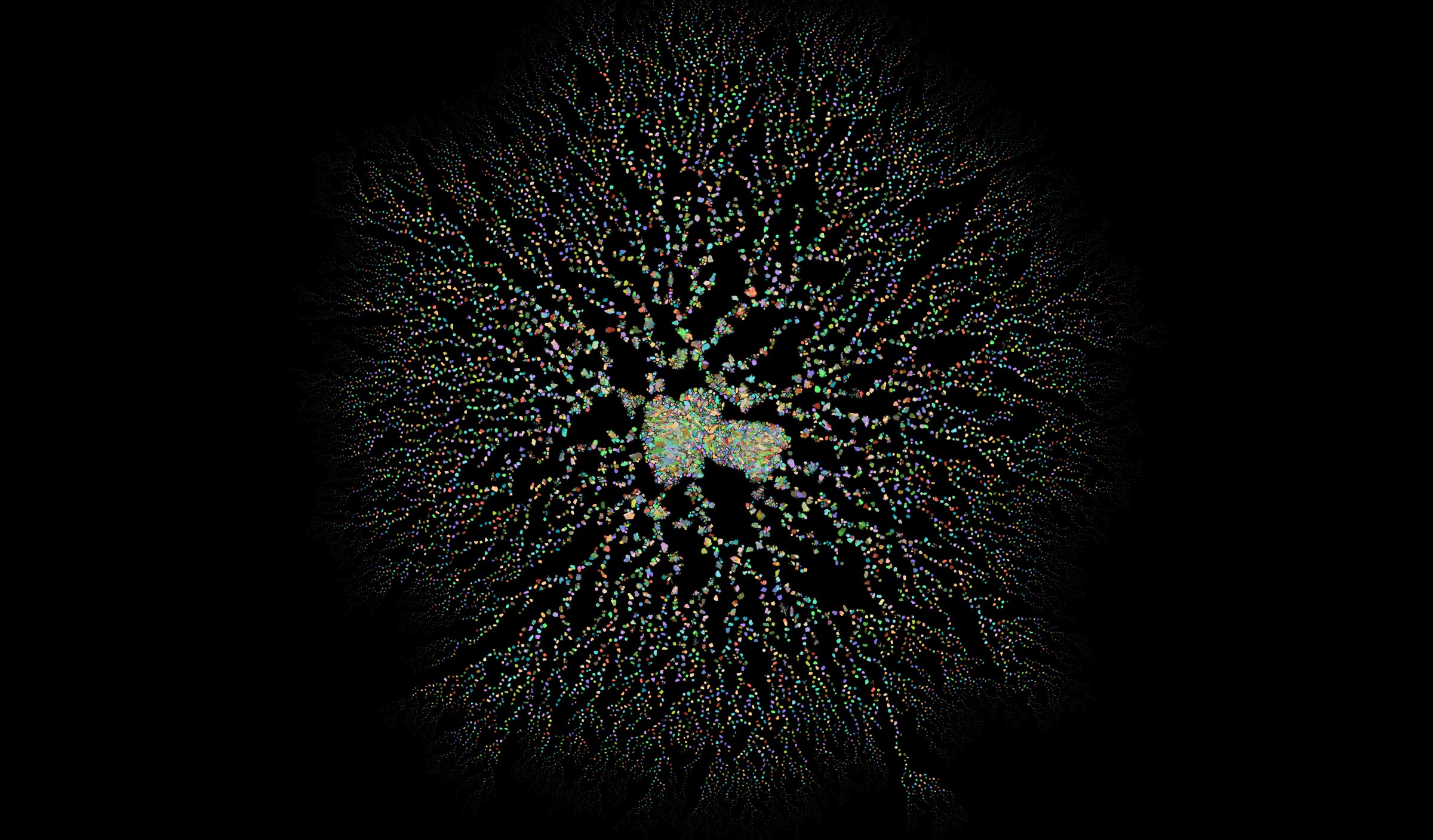
SECOND PLACE
This network was created to visualize evolutionary relatedness of approximately 750,000 genes from 66 different species, ranging from humans to bacteria. Each colored nodule in the network is composed of a set of genes predicted to originate from the same ancestral gene in some ancestral species. When two or more groups can’t be exactly resolved, they are left connected to each other, with the groups of genes at the center being particularly poorly resolved. — Claire McWhite, Cell and Molecular Biology Graduate Student.
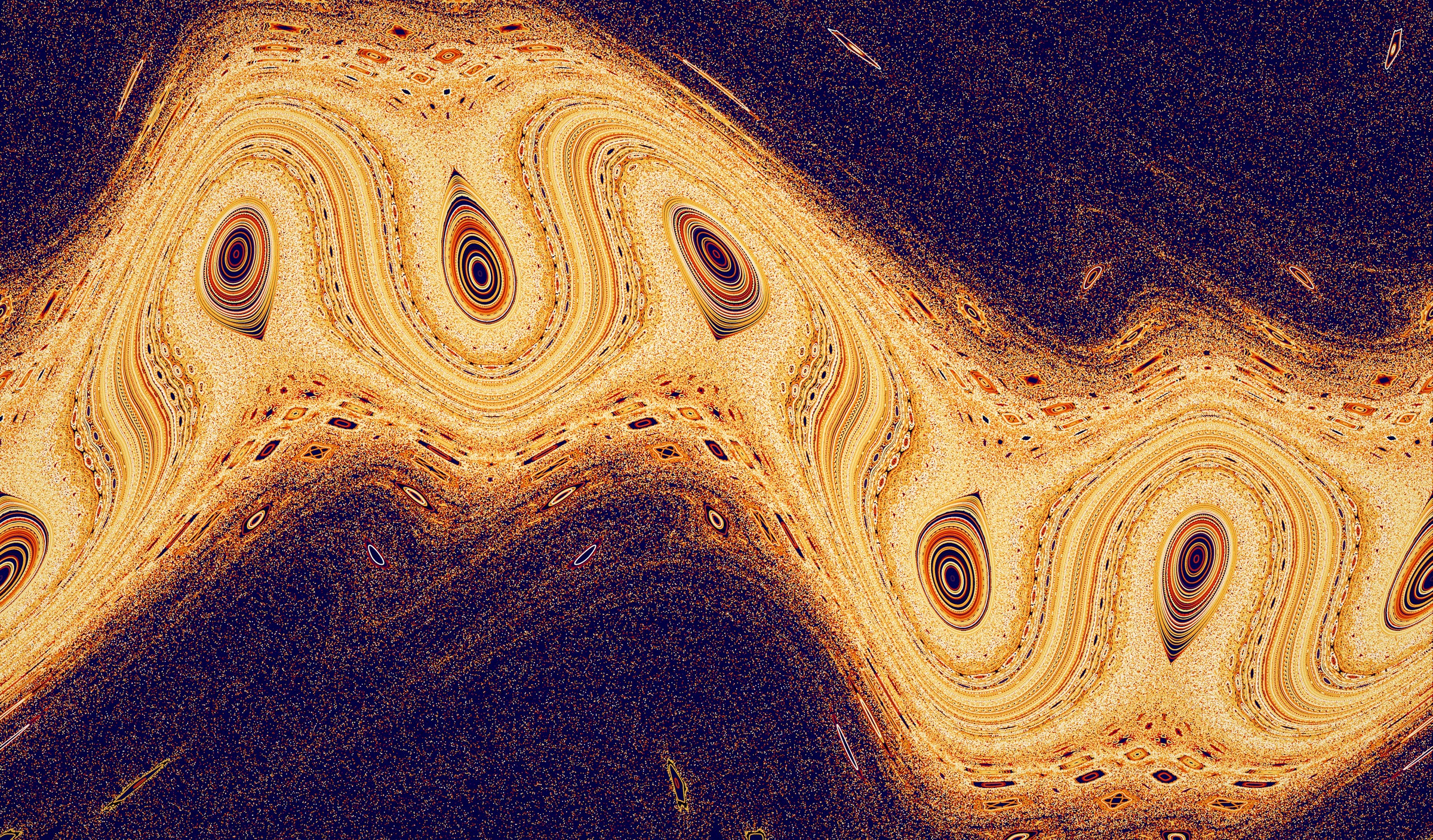
THIRD PLACE
The Standard Nontwist Map is a universal model for describing transport — exchange of mass, energy or momentum — in fluids and gases. It simulates dynamics of particles in various systems and transition to chaotic motion for select map parameters. Regions of high mixing and movement are identified within the chaotic regions on the edge of the map, whereas barriers to transport are identified with orbits in the middle. The study of transport is important for many reasons: for the migration of mid-latitude ozone that replenishes the ozone hole, preventing pollutants in the ocean from washing up on beaches, and preventing the escape of heat and particles in fusion plasma devices. — George Miloshevich, Physics Graduate Student.
HONORABLE MENTIONS
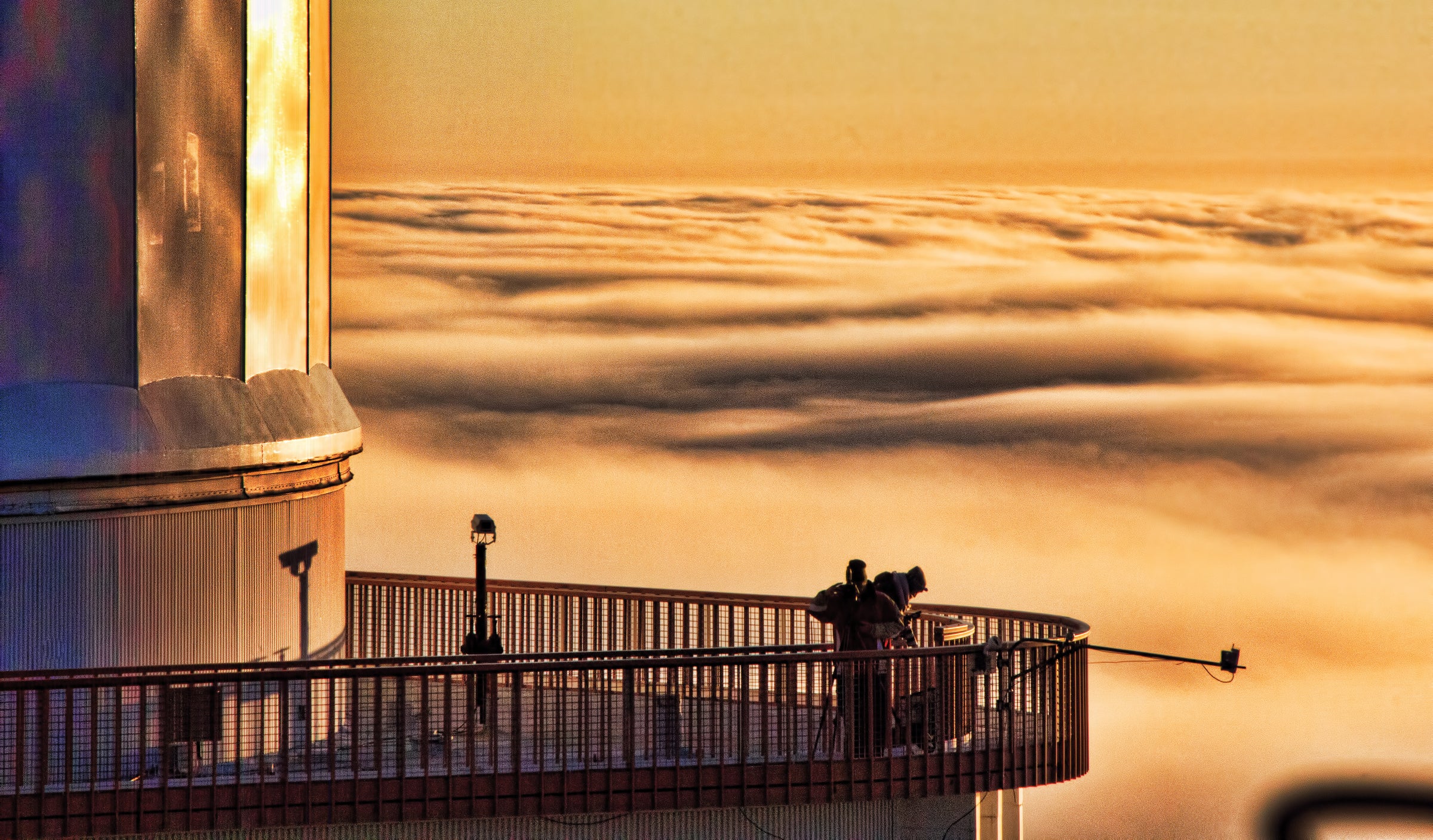
A photograph of Ethan Tweedie and Renee Hoogs, commercial photographers on a shoot for McDonald Observatory, atop the 2.7m Harlan J. Smith telescope dome structure. Photographed from the top of the 2.1m Otto Struve Telescope Dome, with an inversion layer forming a sea of clouds below the Mt. Locke summit. — Coyne Gibson, Manager of Observing Support at McDonald Observatory.
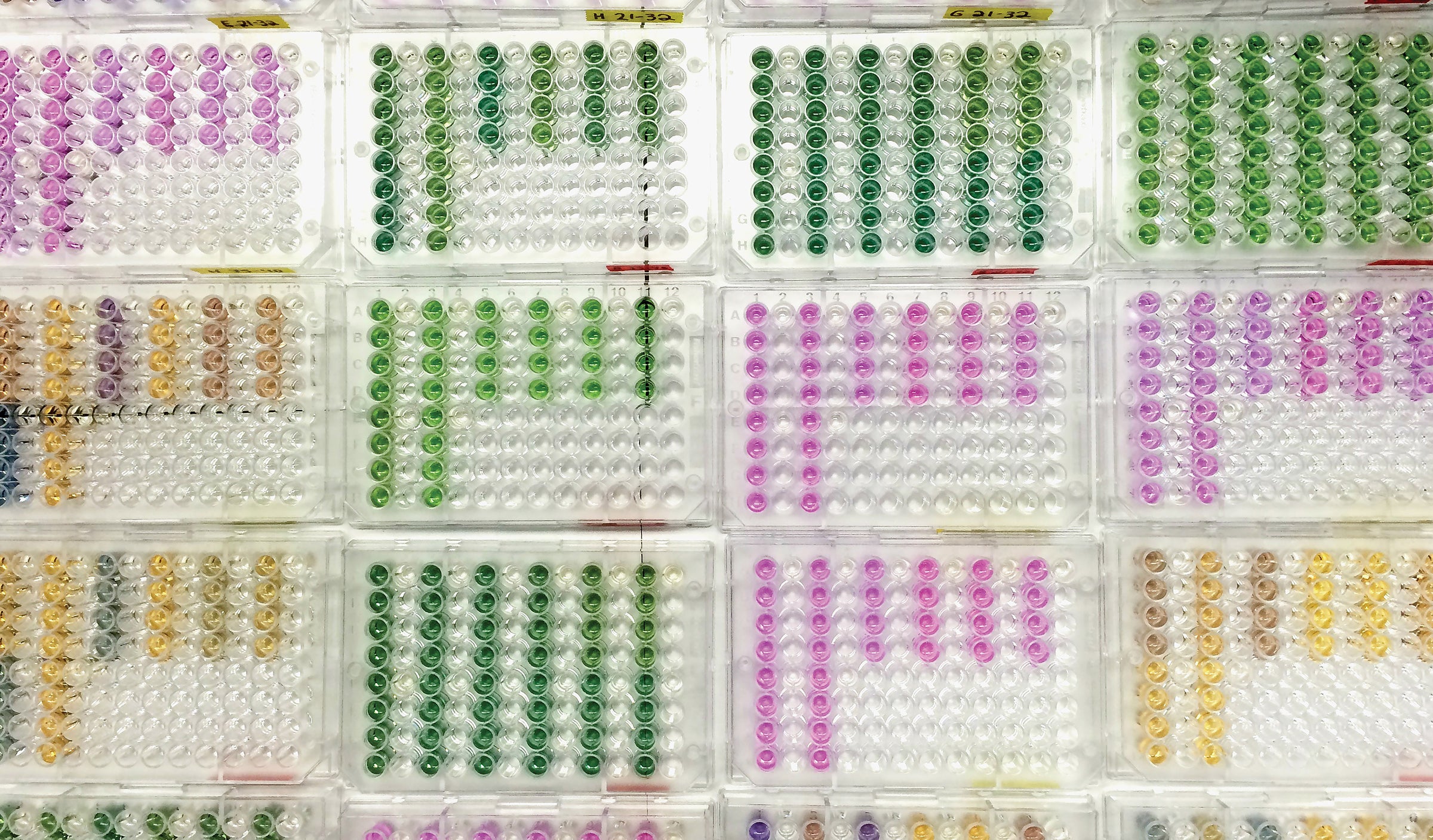
Cachaça is a popular alcoholic beverage in Brazil. These drinks have mixtures of chemical compounds known as tannins that depend on what woods they were soaked with during the production process. Taking advantage of the structure of these molecules, Freshman Research Initiative students in the Supramolecular Sensors Stream created an array of sensing ensembles in an effort to differentiate between cachaça “varietals.” — Elise LeBovidge, Undergraduate Chemistry Student.
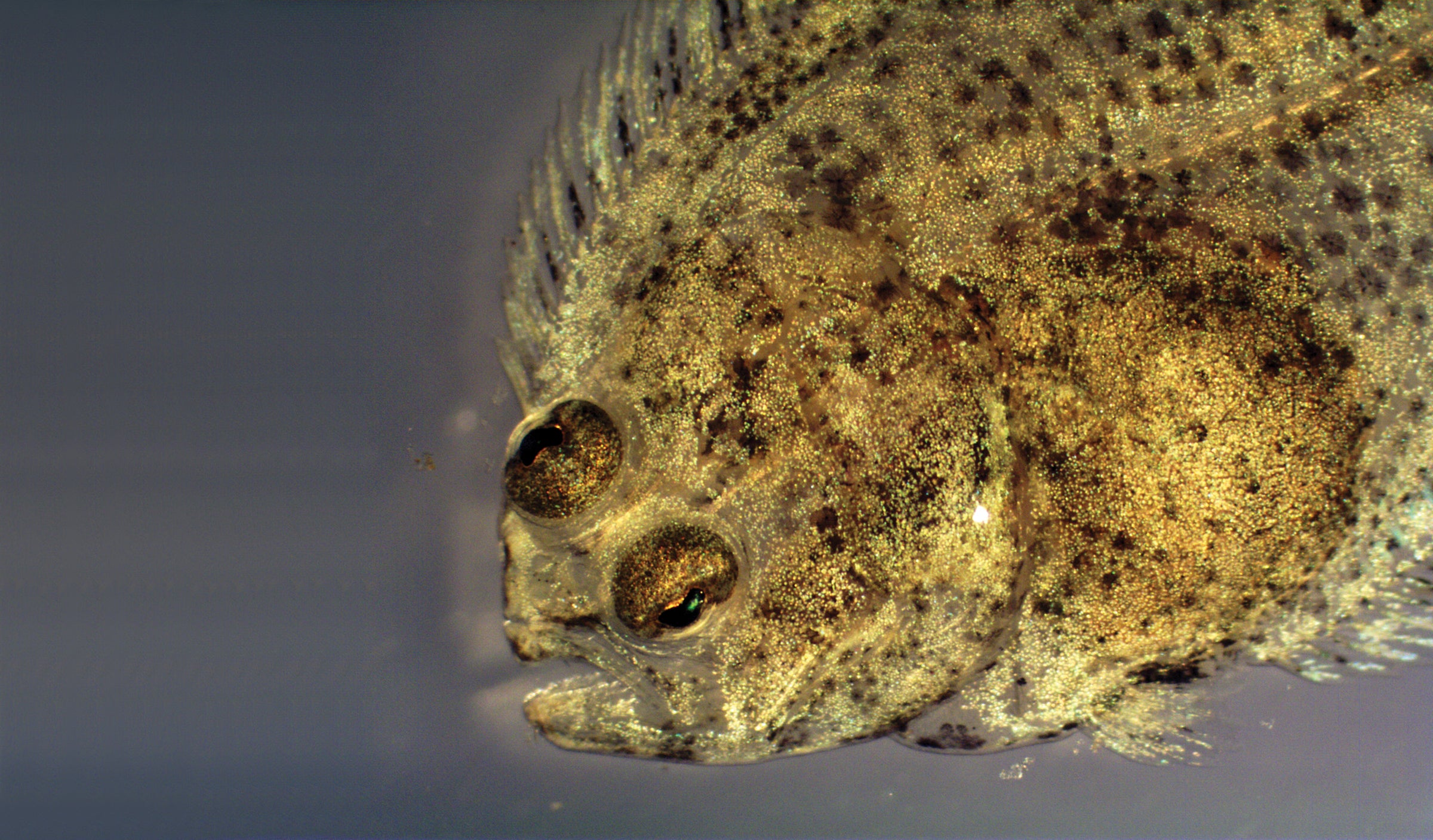
A late-stage metamorphosing Southern flounder, Paralichthys lethostigma, one of many flatfish species, was photographed at 45 days post-hatching when it is just over 1 centimeter in length. Flatfishes undergo a physiologically stressful metamorphosis in their transition from the larval to juvenile life stage. Larval flatfishes are born symmetrical, with one eye on either side of their skull, but during metamorphosis, one eye migrates for multiple days over the top of the head. Eventually both eyes reside permanently on one side of the body while the flatfish lays on its side on the sediment. The first successful spawning of Southern flounder for aquaculture took place at the University of Texas Marine Science Institute’s Fisheries and Mariculture Lab in Port Aransas in 1978 under Dr. Connie Arnold. Today, research at UTSMI FAML is looking into the effects of Southern flounder broodstock nutrition on offspring quality to aid in stock enhancement efforts along the Texas coast. — Corinne Burns, Marine Science Graduate Student.
FACEBOOK FAVORITE
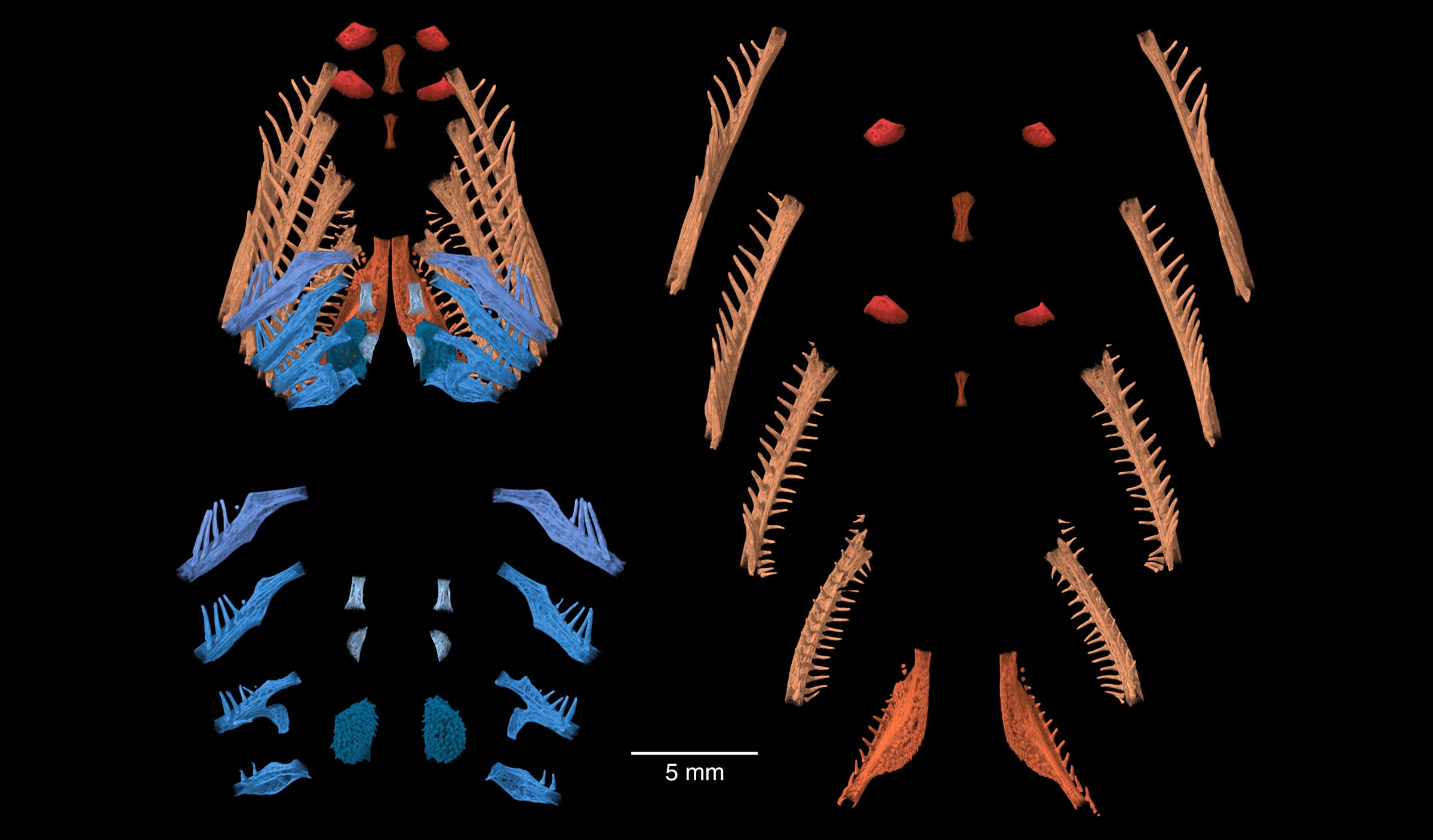
Strange fish live below San Antonio in the biologically diverse, deep pool of the Edwards Aquifer, which provides that city most of its water. One of these fish has a scientific name that starts with “Satan” and is commonly called the eyeless widemouth blind catfish (Satan eurystomus). Its habitat is inaccessible to humans, so specimens of the fish, all collected over 30 years ago, are precious and carefully guarded. A high resolution 3D Computed Tomography scan allowed scientists to digitally extract and analyze details of its skeleton, furthering the knowledge of its evolution, ecology and basic biology. This illustration helps scientists understand how the delicate bones, supporting the species’ gills, function together with its oral jaws for capturing and swallowing prey in the total darkness where it lives. The image was created with a team effort. Dean Hendrickson x-rayed all known specimens, selected the best ossified (Smithsonian specimen USNM 195830), and had it scanned in UT’s Digital Morphology lab. Kyle Luckenbill, Academy of Natural Sciences, Philadelphia, digitally processed the scan data. — Dean Hendrickson, PhD, Curator of Ichthyology in the Department of Integrative Biology.