New Model Reveals How Chromosomes Get Packed Up
The first theoretical model of condensin, a molecular machine involved in packing and unpacking chromosomes, accurately reproduces all known experiments with just two parameters.
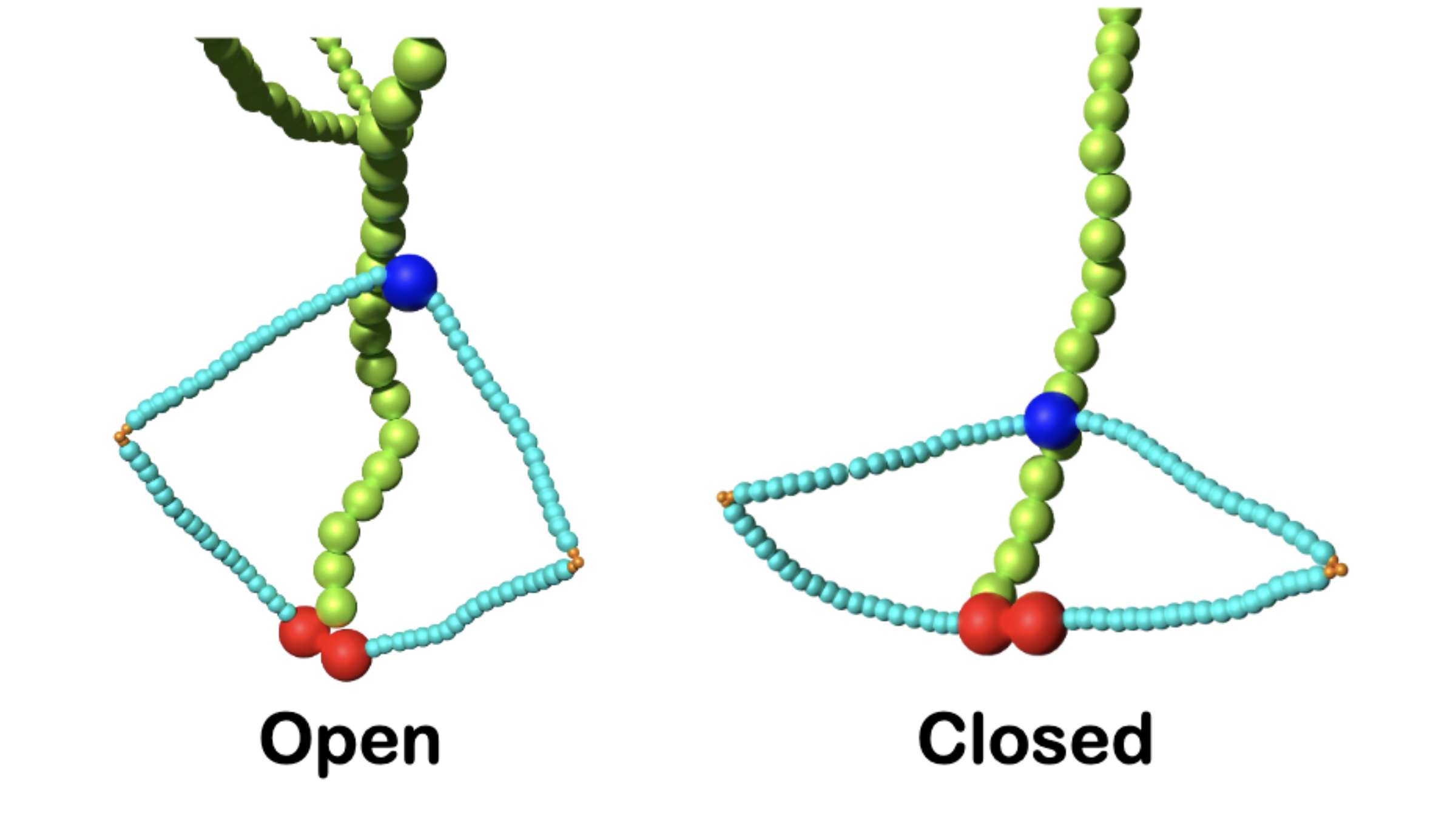
To scrunch a chromosome (green), a condensin molecule opens and closes like a pair of fingers (light blue) connected by a hinge (dark blue).
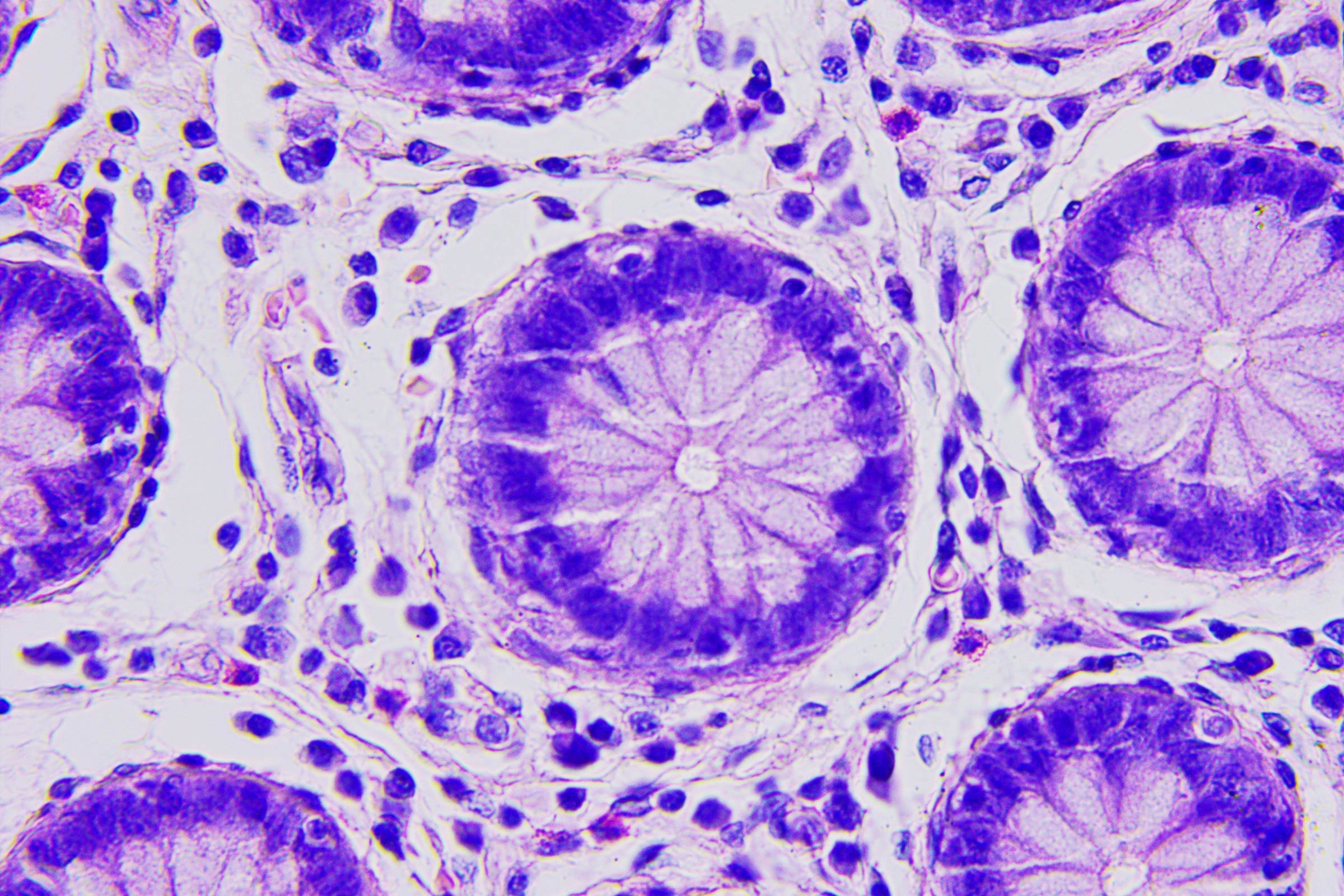
One of the most astounding feats of nature is happening right now in cells throughout your body: noodle-like molecules called chromosomes, which carry part of your genetic blueprints and are about two inches (5 centimeters) long when fully stretched out, get stuffed into the cell's nucleus, which is at least 5,000 times smaller, with plenty of room for a bunch of other chromosomes.
This process repeats every time a cell prepares to divide, which for most cells is about once or twice a day.
For decades, scientists have puzzled over how this packing happens.
Now a team of researchers from The University of Texas at Austin has built the first theoretical model of condensin, a molecular machine involved in packing and unpacking chromosomes, that accurately reproduces all known experiments with just two parameters.
The work, led by professor of chemistry and physics Devarajan (Dave) Thirumalai and Ph.D. candidate Ryota Takaki, is described in a paper published today in the journal Nature Communications.
"I'm really happy to create such a simple equation that actually fits the experiments," Takaki said.
Click to download video: https://www.science.org/doi/suppl/10.1126/science.aar7831/suppl_file/aar7831s4.mp4
In a 2018 paper in Science, researchers from Delft University described how they tethered two ends of a DNA molecule to a surface, stained it with a glowing dye, and added condensin. In this video, you can see a loop of DNA grow (get brighter) as condensin scrunches up the strand of DNA.
Earlier experiments showed that condensin reels in the chromosome, scrunching sections up into thousands of loops, causing the chromosome to look less like a long, straight noodle and more like a dense ball of spaghetti. Condensin looks a bit like two fingers joined together by a hinge at the base. To reel in a section of chromosome, the fingers pull inward toward the base.
The new model built by Takaki, which accurately reproduces the speed of loop formation from those earlier experiments on chromosomes, uses just two parameters: the distance between the finger tips and base of condensin as it opens and closes, and the rate that condensin consumes energy. Scientists often describe theories that are both simple and powerful as being beautiful.
"As theorists, we're constantly looking for beautiful theories," said Thirumalai, the Marvin K. Collie-Welch Regents Chair in Chemistry and senior author of the paper. "Sometimes it works, sometimes it doesn't. In this case, it does."
Chromosomes are a vital element in every cell; they contain the genetic information for building all the proteins in the body and maintaining all the normal functions of life. When condensin doesn't perform its role in packing and unpacking chromosomes correctly, the organism usually dies early in development. Meanwhile, some mutations can lead to various forms of cancer, most commonly T-cell lymphoma.
"This packing and unpacking has to happen for many, many cycles during the life of a cell," Thirumalai said. "If it doesn't work correctly, that has big impacts downstream. It's like when your computer's hard drive crashes, you're in big trouble. Fortunately, we have systems in the cell that usually keep that from happening."
The paper's other authors are graduate student Atreya Dey and postdoctoral researcher Guang Shi.